Data file must be an archive in .zip or .tar format and contain valid PDB files of conformational
ensembles. PDB files can be generated with any program. For our studies we generated models
using all-atom Monte Carlo simulations as implemented in Rosetta.
Example PDB archive file (*.zip) can be found
here
Intensity profiles can be either simulated "on-the-fly" with
Pepsi-SAXS method
or uploaded from your drive. If you want to use the latter option the file must have numeric
values and contain n x m columns (n = number of q points, m = number of models) separated with the space.
Profiles have to be simulated for the same q points as in experimental file and therefor their number
must be the same in both files (experimental and simulated profiles).
Example of simulated profiles for 10 q points and 5 structural models can be found
here
Jensen-Shannon is a useful metric to measure the uncertainty of ensembles developed by (Fisher et al*).
The expectation value of the Jensen-Shannon divergence relative to the optimal weights over
the posterior distribution can be defined as:
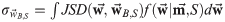 where:
where:
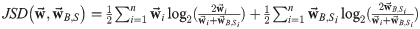 and ranges between 0 and 1 for two maximally identical and different vectors.
and ranges between 0 and 1 for two maximally identical and different vectors.
*Fisher CK, Ullman O, Stultz CM. Pacific Symposium on Biocomputing Pacific Symposium on Biocomputing. 2012:82–93. Epub 2011/12/17. pmid:22174265
Model evidence or marginal likelihood is widely used in Bayesian model comparison
and provides an automatic Occam’s razor effect by balancing between fit to data
and model complexity, thereby providing a rigorous approach to combat overfitting.
However, ,odel evidence is a multidimensional integral that can be very difficult to evaluate.
We use TIStan* package that implements
adaptively-annealed thermodynamic integration for model evidence estimation.
This package makes use of PyStan's implementation of the No U-Turn Sampler for refreshing
the sample population at each inverse temperature increment.
*Henderson, R.W.; Goggans, P.M. TI-Stan: Model Comparison Using Thermodynamic Integration and HMC. Entropy 2019, 21, 1161.
Correlation Map (CorMap) is a measure for assessing differences
between one-dimensional spectra independently of explicit error estimates,
using only data point correlations.* CorMap identifies the longest stretch (C) of data points that
lie on one side of the model profile and provides a probability (P) for that occurrence
given the number of points (n) in the data set.
We use corrmap implementation from freesas package
*Franke, D., Jeffries, C. & Svergun, D.
Correlation Map, a goodness-of-fit test for one-dimensional X-ray scattering spectra.
Nat Methods 12, 419–422 (2015). https://doi.org/10.1038/nmeth.3358
Calmodulin (CaM) is a two-domain protein system connected by the flexible linker. We will analyze SAXS data set of CaM obtained using in-line SEC (size exclusion chromatography) at the Australian Synchrotron* Library of structurally and energetically reasonable conformers of CaM was generated using Rosetta macromolecular modeling package where torsion angles in the linker segment were sampled in a Monte Carlo simulation followed by an all atom energy refinement of the linker segment (residues 77-81).
Defining input and running analysis:
- Upload SAXS data and pdb files (login required).
- Click on button and input analysis name.
- Weight cutoff controls number of models discarded after each iterations (keep the default value) and may influence final number of models selected for ensemble.
- Number of iterations controls STAN simulations and 2000 is usually good number (keep the default value)
- CLick on Analyze button. This will start analysis and you will receive email when it is finished
- You will see if job is running on your profile page (under Analyses).
- Once job's finished you can inspect results by clicking on
Results:
The process is stochastic, so results may change a bit from run to run.
Nevertheless in this case, we expect three structures to be selected and overall good fit
to the data (χ2) ~ 0.78, Jensen-Shanon divergence ~ 0.05 and Model Evidence of approx. -800.
*Trewhella J, Duff AP, Durand D, Gabel F, Guss JM, Hendrickson WA, et al. 2017 publication guidelines for structural modelling of small-angle scattering data from biomolecules in solution: an update. Acta crystallographica Section D, Structural biology. 2017;73(Pt 9):710–28. Epub 2017/09/07. pmid:28876235.
Once analsysis is finished one can download results by clicking on
Downloaded archive contains several files:
- vbi_output.txt Full output from variational bayesian inference including all weights per iteration
- vbi_output.txt.log Output summary from variational bayesian inference.
- vbi_output.txt.dat Fit file with q, I(q), Isim(q), err(q) from variational bayesian inference.
- cbi_output.txt Output summary from complete bayesian inference.
- cbi_output.txt.fit Fit file with q, I(q), Isim(q), err(q) from complete bayesian inference.
- saved_samples.txt Weights samples generated by pystan
- stan_weight_*.png Weight distribution plot for each PDB file
- *.pdb PDB files selected for final ensemble
- ensemble.pdb Combined pdb file with selected models
- fit.png Intensity plot with selected enesemble fitted to pdb file